没有合适的资源?快使用搜索试试~ 我知道了~
Dermatologist-level classification of skin cancer(这可是nature呀)1
需积分: 0 0 下载量 27 浏览量
2022-08-03
15:20:58
上传
评论
收藏 2.81MB PDF 举报
温馨提示

试读
12页
Dermatologist-level classification of skin cancer(这可是nature呀)1
资源详情
资源评论
资源推荐

2 FEBRUARY 2017 | VOL 542 | NATURE | 115
LETTER
doi:10.1038/nature21056
Dermatologist-level classification of skin cancer
with deep neural networks
Andre Esteva
1
*, Brett Kuprel
1
*, Roberto A. Novoa
2,3
, Justin Ko
2
, Susan M. Swetter
2,4
, Helen M. Blau
5
& Sebastian Thrun
6
Skin cancer, the most common human malignancy
1–3
, is primarily
diagnosed visually, beginning with an initial clinical screening
and followed potentially by dermoscopic analysis, a biopsy and
histopathological examination. Automated classification of skin
lesions using images is a challenging task owing to the fine-grained
variability in the appearance of skin lesions. Deep convolutional
neural networks (CNNs)
4,5
show potential for general and highly
variable tasks across many fine-grained object categories
6–11
.
Here we demonstrate classification of skin lesions using a single
CNN, trained end-to-end from images directly, using only pixels
and disease labels as inputs. We train a CNN using a dataset of
129,450clinical images—two orders of magnitude larger than
previous datasets
12
—consisting of 2,032different diseases. We
test its performance against 21board-certified dermatologists on
biopsy-proven clinical images with two critical binary classification
use cases: keratinocyte carcinomas versus benign seborrheic
keratoses; and malignant melanomas versus benign nevi. The first
case represents the identification of the most common cancers, the
second represents the identification of the deadliest skin cancer.
The CNN achieves performance on par with all tested experts
across both tasks, demonstrating an artificial intelligence capable
of classifying skin cancer with a level of competence comparable to
dermatologists. Outfitted with deep neural networks, mobile devices
can potentially extend the reach of dermatologists outside of the
clinic. It is projected that 6.3billion smartphone subscriptions will
exist by the year 2021 (ref. 13) and can therefore potentially provide
low-cost universal access to vital diagnostic care.
There are 5.4million new cases of skin cancer in the United States
2
every year. One in five Americans will be diagnosed with a cutaneous
malignancy in their lifetime. Although melanomas represent fewer than
5% of all skin cancers in the United States, they account for approxi-
mately 75% of all skin-cancer-related deaths, and are responsible for
over 10,000deaths annually in the United States alone. Early detection
is critical, as the estimated 5-year survival rate for melanoma drops
from over 99% if detected in its earliest stages to about 14% if detected
in its latest stages. We developed a computational method which may
allow medical practitioners and patients to proactively track skin
lesions and detect cancer earlier. By creating a novel disease taxonomy,
and a disease-partitioning algorithm that maps individual diseases into
training classes, we are able to build a deep learning system for auto-
mated dermatology.
Previous work in dermatological computer-aided classification
12,14,15
has lacked the generalization capability of medical practitioners
owing to insufficient data and a focus on standardized tasks such as
dermoscopy
16–18
and histological image classification
19–22
. Dermoscopy
images are acquired via a specialized instrument and histological
images are acquired via invasive biopsy and microscopy; whereby
both modalities yield highly standardized images. Photographic
images (for example, smartphone images) exhibit variability in factors
such as zoom, angle and lighting, making classification substantially
more challenging
23,24
. We overcome this challenge by using a data-
driven approach—1.41million pre-training and training images
make classification robust to photographic variability. Many previous
techniques require extensive preprocessing, lesion segmentation and
extraction of domain-specific visual features before classification. By
contrast, our system requires no hand-crafted features; it is trained
end-to-end directly from image labels and raw pixels, with a single
network for both photographic and dermoscopic images. The existing
body of work uses small datasets of typically less than a thousand
images of skin lesions
16,18,19
, which, as a result, do not generalize well
to new images. We demonstrate generalizable classification with a new
dermatologist-labelled dataset of 129,450clinical images, including
3,374dermoscopy images.
Deep learning algorithms, powered by advances in computation
and very large datasets
25
, have recently been shown to exceed human
performance in visual tasks such as playing Atari games
26
, strategic
board games like Go
27
and object recognition
6
. In this paper we
outline the development of a CNN that matches the performance of
dermatologists at three key diagnostic tasks: melanoma classification,
melanoma classification using dermoscopy and carcinoma
classification. We restrict the comparisons to image-based classification.
We utilize a GoogleNet Inception v3 CNN architecture
9
that was pre-
trained on approximately 1.28million images (1,000object categories)
from the 2014 ImageNet Large Scale Visual Recognition Challenge
6
,
and train it on our dataset using transfer learning
28
. Figure 1 shows the
working system. The CNN is trained using 757disease classes. Our
dataset is composed of dermatologist-labelled images organized in a
tree-structured taxonomy of 2,032diseases, in which the individual
diseases form the leaf nodes. The images come from 18different
clinician-curated, open-access online repositories, as well as from
clinical data from Stanford University Medical Center. Figure 2a shows
a subset of the full taxonomy, which has been organized clinically and
visually by medical experts. We split our dataset into 127,463training
and validation images and 1,942biopsy-labelled test images.
To take advantage of fine-grained information contained within the
taxonomy structure, we develop an algorithm (Extended Data Table 1)
to partition diseases into fine-grained training classes (for example,
amelanotic melanoma and acrolentiginous melanoma). During
inference, the CNN outputs a probability distribution over these fine
classes. To recover the probabilities for coarser-level classes of interest
(for example, melanoma) we sum the probabilities of their descendants
(see Methods and Extended Data Fig. 1 for more details).
We validate the effectiveness of the algorithm in two ways, using
nine-fold cross-validation. First, we validate the algorithm using a
three-class disease partition—the first-level nodes of the taxonomy,
which represent benign lesions, malignant lesions and non-neoplastic
1
Department of Electrical Engineering, Stanford University, Stanford, California, USA.
2
Department of Dermatology, Stanford University, Stanford, California, USA.
3
Department of Pathology,
Stanford University, Stanford, California, USA.
4
Dermatology Service, Veterans Affairs Palo Alto Health Care System, Palo Alto, California, USA.
5
Baxter Laboratory for Stem Cell Biology, Department
of Microbiology and Immunology, Institute for Stem CellBiology and RegenerativeMedicine, Stanford University, Stanford, California, USA.
6
Department of Computer Science, Stanford University,
Stanford, California, USA.
*These authors contributed equally to this work.
© 2017 Macmillan Publishers Limited, part of Springer Nature. All rights reserved.

116 | NATURE | VOL 542 | 2 FEBRUARY 2017
Letter
reSeArCH
lesions. In this task, the CNN achieves 72.1 ± 0.9% (mean ± s.d.) overall
accuracy (the average of individual inference class accuracies) and two
dermatologists attain 65.56% and 66.0% accuracy on a subset of the
validation set. Second, we validate the algorithm using a nine-class
disease partition—the second-level nodes—so that the diseases of
each class have similar medical treatment plans. The CNN achieves
55.4 ± 1.7% overall accuracy whereas the same two dermatologists
attain 53.3% and 55.0% accuracy. A CNN trained on a finer disease
partition performs better than one trained directly on three or nine
classes (see Extended Data Table 2), demonstrating the effectiveness
of our partitioning algorithm. Because images of the validation set are
labelled by dermatologists, but not necessarily confirmed by biopsy,
this metric is inconclusive, and instead shows that the CNN is learning
relevant information.
To conclusively validate the algorithm, we tested, using only
biopsy-proven images on medically important use cases, whether
the algorithm and dermatologists could distinguish malignant versus
benign lesions of epidermal (keratinocyte carcinoma compared to
benign seborrheic keratosis) or melanocytic (malignant melanoma
compared to benign nevus) origin. For melanocytic lesions, we show
two trials, one using standard images and the other using dermoscopy
images, which reflect the two steps that a dermatologist might carry out
to obtain a clinical impression. The same CNN is used for all three tasks.
Figure 2b shows a few example images, demonstrating the difficulty in
distinguishing between malignant and benign lesions, which share many
visual features. Our comparison metrics are sensitivity and specificity:
=sensitivity
true positive
positive
=specificity
true negative
negative
where ‘true positive’ is the number of correctly predicted malignant
lesions, ‘positive’ is the number of malignant lesions shown, ‘true neg-
ative’ is the number of correctly predicted benign lesions, and ‘neg-
ative’ is the number of benign lesions shown. When a test set is fed
through the CNN, it outputs a probability, P, of malignancy, per image.
We can compute the sensitivity and specificity of these probabilities
Acral-lentiginous melanoma
Amelanotic melanoma
Lentigo melanoma
…
Blue nevus
Halo nevus
Mongolian spot
…
Training classes (757)Deep convolutional neural network (Inception v3)
Inference classes (varies by task)
92% malignant melanocytic lesion
8% benign melanocytic lesion
Skin lesion image
Convolution
AvgPool
MaxPool
Concat
Dropout
Fully connected
Softmax
Figure 1 | Deep CNN layout. Our classification technique is a
deep CNN. Data flow is from left to right: an image of a skin lesion
(for example, melanoma) is sequentially warped into a probability
distribution over clinical classes of skin disease using Google Inception
v3 CNN architecture pretrained on the ImageNet dataset (1.28million
images over 1,000generic object classes) and fine-tuned on our own
dataset of 129,450skin lesions comprising 2,032different diseases.
The 757training classes are defined using a novel taxonomy of skin disease
and a partitioning algorithm that maps diseases into training classes
(for example, acrolentiginous melanoma, amelanotic melanoma, lentigo
melanoma). Inference classes are more general and are composed of one
or more training classes (for example, malignant melanocytic lesions—the
class of melanomas). The probability of an inference class is calculated by
summing the probabilities of the training classes according to taxonomy
structure (see Methods). Inception v3 CNN architecture reprinted
from https://research.googleblog.com/2016/03/train-your-own-image-
classifier-with.html
ba
Epidermal lesions
Benign
Malignant
Melanocytic lesions Melanocytic lesions (dermoscopy)
Skin disease
Benign
Melanocytic
Café au
lait spot
Solar
lentigo
Epidermal
Seborrhoeic
keratosis
Milia
Dermal
Cyst
Non-
neoplastic
Acne
Rosacea
Abrasion
Stevens-
Johnson
syndrome
Tuberous
sclerosis
Malignant
Epidermal
Basal cell
carcinoma
Squamous
cell
carcinoma
Dermal
Merkel cell
carcinoma
Angiosarcoma
T-cell
B-cell
Genodermatosis
Congenital
dyskeratosis
Bullous
pemphigoid
Cutaneous
lymphoma
Melanoma
Psoriasis
Fibroma
Lipoma
Inammatory
Atypical
nevus
Figure 2 | A schematic illustration of the taxonomy and example test
set images. a, A subset of the top of the tree-structured taxonomy of skin
disease. The full taxonomy contains 2,032diseases and is organized based
on visual and clinical similarity of diseases. Red indicates malignant,
green indicates benign, and orange indicates conditions that can be either.
Black indicates melanoma. The first two levels of the taxonomy are used in
validation. Testing is restricted to the tasks of b. b, Malignant and benign
example images from two disease classes. These test images highlight the
difficulty of malignant versus benign discernment for the three medically
critical classification tasks we consider: epidermal lesions, melanocytic
lesions and melanocytic lesions visualized with a dermoscope. Example
images reprinted with permission from the Edinburgh Dermofit Library
(https://licensing.eri.ed.ac.uk/i/software/dermofit-image-library.html).
© 2017 Macmillan Publishers Limited, part of Springer Nature. All rights reserved.
2 FEBRUARY 2017 | VOL 542 | NATURE | 117
Letter
reSeArCH
by choosing a threshold probability t and defining the prediction ŷ for
each image as ŷ = P ≥ t. Varying t in the interval 0–1 generates a curve
of sensitivities and specificities that the CNN can achieve.
We compared the direct performance of the CNN and at least
21board-certified dermatologists on epidermal and melanocytic
lesion classification (Fig. 3a). For each image the dermatologists
were asked whether to biopsy/treat the lesion or reassure the patient.
Each red point on the plots represents the sensitivity and specificity
of a single dermatologist. The CNN outperforms any dermatologist
whose sensitivity and specificity point falls below the blue curve of
a
b
0 1
Sensitivity
0
1
Specicity
Melanoma: 130 images
0 1
Sensitivity
0
1
Specicity
Melanoma: 225 images
Algorithm: AUC = 0.96
0 1
Sensitivity
0
1
Specicity
Melanoma: 111 dermoscopy images
0 1
Sensitivity
0
1
Specicity
Carcinoma: 707 images
Algorithm: AUC = 0.96
0 1
Sensitivity
0
1
Specicity
Melanoma: 1,010 dermoscopy image
s
Algorithm: AUC = 0.94
0 1
Sensitivity
0
1
Specicity
Carcinoma: 135 images
Algorithm: AUC = 0.96
Dermatologists (25)
Average dermatologist
Algorithm: AUC = 0.94
Dermatologists (22)
Average dermatologist
Algorithm: AUC = 0.91
Dermatologists (21)
Average dermatologist
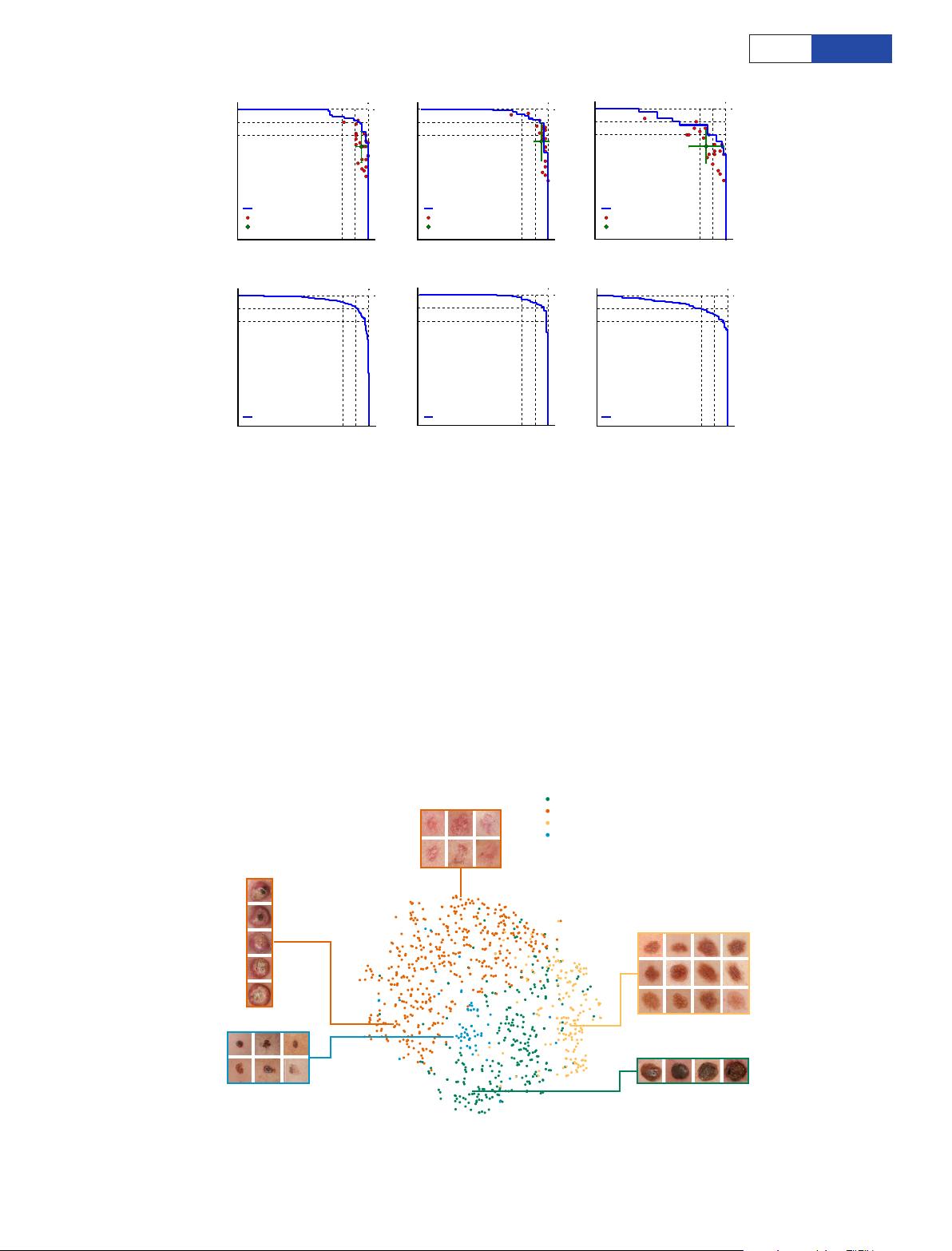
Figure 3 | Skin cancer classification performance of the CNN and
dermatologists. a, The deep learning CNN outperforms the average of
the dermatologists at skin cancer classification using photographic and
dermoscopic images. Our CNN is tested against at least 21dermatologists
at keratinocyte carcinoma and melanoma recognition. For each test,
previously unseen, biopsy-proven images of lesions are displayed, and
dermatologists are asked if they would: biopsy/treat the lesion or reassure
the patient. Sensitivity, the true positive rate, and specificity, the true
negative rate, measure performance. A dermatologist outputs a single
prediction per image and is thus represented by a single red point. The
green points are the average of the dermatologists for each task, with
error bars denoting one standard deviation (calculated from n = 25, 22
and 21tested dermatologists for keratinocyte carcinoma, melanoma
and melanoma under dermoscopy, respectively). The CNN outputs a
malignancy probability P per image. We fix a threshold probability t
such that the prediction ŷ for any image is ŷ = P ≥ t, and the blue curve is
drawn by sweeping t in the interval 0–1. The AUC is the CNN’s measure
of performance, with a maximum value of 1. The CNN achieves superior
performance to a dermatologist if the sensitivity–specificity point of
the dermatologist lies below the blue curve, which most do. Epidermal
test: 65keratinocyte carcinomas and 70benign seborrheic keratoses.
Melanocytic test: 33malignant melanomas and 97benign nevi. A second
melanocytic test using dermoscopic images is displayed for comparison:
71malignant and 40benign. The slight performance decrease reflects
differences in the difficulty of the images tested rather than the diagnostic
accuracies of visual versus dermoscopic examination. b, The deep learning
CNN exhibits reliable cancer classification when tested on a larger dataset.
We tested the CNN on more images to demonstrate robust and reliable
cancer classification. The CNN’s curves are smoother owing to the larger
test set.
Squamous cell ca
rcinomas
Basal cell carcinomas
Nevi
Melanomas
Seborrhoeic keratoses
Epidermal benign
Epidermal malignant
Melanocytic benign
Melanocytic malignant
Figure 4 | t-SNE visualization of the last hidden layer representations
in the CNN for four disease classes. Here we show the CNN’s internal
representation of four important disease classes by applying t-SNE,
a method for visualizing high-dimensional data, to the last hidden layer
representation in the CNN of the biopsy-proven photographic test sets
(932images). Coloured point clouds represent the different disease
categories, showing how the algorithm clusters the diseases. Insets show
images corresponding to various points. Images reprinted with permission
from the Edinburgh Dermofit Library (https://licensing.eri.ed.ac.uk/i/
software/dermofit-image-library.html).
© 2017 Macmillan Publishers Limited, part of Springer Nature. All rights reserved.
剩余11页未读,继续阅读
被要求改名字
- 粉丝: 26
- 资源: 315
上传资源 快速赚钱
 我的内容管理
展开
我的内容管理
展开
 我的资源
快来上传第一个资源
我的资源
快来上传第一个资源
 我的收益 登录查看自己的收益
我的收益 登录查看自己的收益 我的积分
登录查看自己的积分
我的积分
登录查看自己的积分
 我的C币
登录后查看C币余额
我的C币
登录后查看C币余额
 我的收藏
我的收藏  我的下载
我的下载  下载帮助
下载帮助

 前往需求广场,查看用户热搜
前往需求广场,查看用户热搜安全验证
文档复制为VIP权益,开通VIP直接复制
 信息提交成功
信息提交成功

评论0