# pyHH
**pyHH is a simple Python implementation of the Hodgkin-Huxley spiking neuron model.** pyHH simulates conductances and calculates membrane voltage at discrete time points without requiring a differential equation solver. [HHSharp](https://github.com/swharden/HHSharp) is a similar project written in C#.

### Minimal Code Example
A full Hodgkin-Huxley spiking neuron model and simulation was created in fewer than 100 lines of Python ([dev/concept4.py](dev/concept4.py)). Unlike other code examples on the internet, this implementation is object-oriented and Pythonic. When run, it produces the image above.
## Python Package
The `pyhh` package includes Hodgkin-Huxley models and additional tools to organize simulation data.
### Simulation Steps
1. Create a model cell and customize its properties if desired
2. Create a stimulus waveform (a numpy array)
3. Create a simulation by giving it model the waveform you created
4. Plot various properties of the stimulation
### Example Usage
```python
# customize a neuron model if desired
model = pyhh.HHModel()
model.gNa = 100 # typically 120
model.gK = 5 # typically 36
model.EK = -35 # typically -12
# customize a stimulus waveform
stim = np.zeros(20000)
stim[7000:13000] = 50 # add a square pulse
# simulate the model cell using the custom waveform
sim = pyhh.Simulation(model)
sim.Run(stimulusWaveform=stim, stepSizeMs=0.01)
```
```python
# plot the results with MatPlotLib
plt.figure(figsize=(10, 8))
ax1 = plt.subplot(411)
ax1.plot(sim.times, sim.Vm - 70, color='b')
ax1.set_ylabel("Potential (mV)")
ax1.set_title("Hodgkin-Huxley Spiking Neuron Model", fontSize=16)
ax2 = plt.subplot(412)
ax2.plot(sim.times, stim, color='r')
ax2.set_ylabel("Stimulation (µA/cm²)")
ax3 = plt.subplot(413, sharex=ax1)
ax3.plot(sim.times, sim.StateH, label='h')
ax3.plot(sim.times, sim.StateM, label='m')
ax3.plot(sim.times, sim.StateN, label='n')
ax3.set_ylabel("Activation (frac)")
ax3.legend()
ax4 = plt.subplot(414, sharex=ax1)
ax4.plot(sim.times, sim.INa, label='VGSC')
ax4.plot(sim.times, sim.IK, label='VGKC')
ax4.plot(sim.times, sim.IKleak, label='KLeak')
ax4.set_ylabel("Current (µA/cm²)")
ax4.set_xlabel("Simulation Time (milliseconds)")
ax4.legend()
plt.tight_layout()
plt.savefig("tests/demo.png")
plt.show()
```

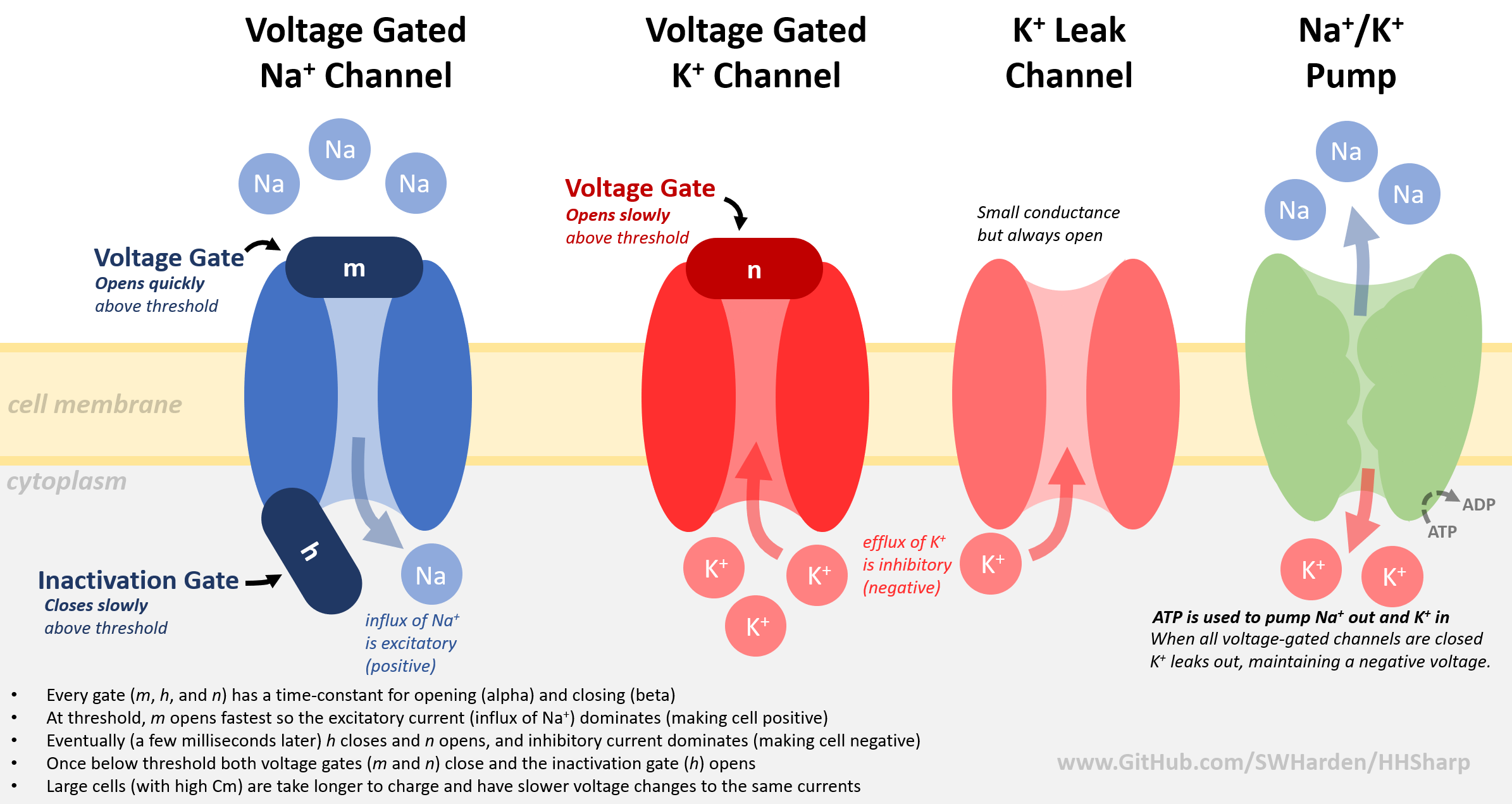
## Theory
Visit https://github.com/swharden/HHSharp for code concepts and simulation notes. Although the language is different, the biology and implementation is the same.
### Additional Resources
* [Hodgkin and Huxley, 1952](https://www.ncbi.nlm.nih.gov/pmc/articles/PMC1392413/pdf/jphysiol01442-0106.pdf) (the original manuscript)
* [The Hodgkin-Huxley Mode](http://www.genesis-sim.org/GENESIS/iBoG/iBoGpdf/chapt4.pdf) (The GENESIS Simulator, Chapter 4)
* Wikipedia: [Hodgkin–Huxley model](https://en.wikipedia.org/wiki/Hodgkin%E2%80%93Huxley_model)
* [Hodgkin-Huxley spiking neuron model in Python](https://www.bonaccorso.eu/2017/08/19/hodgkin-huxley-spiking-neuron-model-python/) by Giuseppe Bonaccorso - a HH model which uses the [`scipy.integrate.odeint` ordinary differential equation solver](https://docs.scipy.org/doc/scipy/reference/generated/scipy.integrate.odeint.html)
* [Introduction to Computational Modeling: Hodgkin-Huxley Model](http://andysbrainblog.blogspot.com/2013/10/introduction-to-computational-modeling.html) by Andrew Jahn - a commentary of the HH model with matlab code which discretely simulates conductances
* [NeuroML Hodgkin Huxley Tutorials](https://github.com/swharden/hodgkin_huxley_tutorial)
* [Summary of the Hodgkin-Huxley model](http://ecee.colorado.edu/~ecen4831/HHsumWWW/HHsum.html) by Dave Beeman
没有合适的资源?快使用搜索试试~ 我知道了~
matlab微分方程代码-pyHH:Hodgkin-Huxley尖峰神经元模型的简单Python实现

共16个文件
py:8个
png:5个
gitignore:1个
温馨提示
matlab微分方程代码pyHH pyHH是Hodgkin-Huxley峰值神经元模型的简单Python实现。 pyHH可以模拟电导并计算离散时间点的膜电压,而无需使用微分方程求解器。 是一个用C#编写的类似项目。 最小代码示例 在不到100行的Python()中创建了完整的Hodgkin-Huxley峰值神经元模型和仿真。 与Internet上的其他代码示例不同,此实现是面向对象的和Pythonic的。 运行时,将产生上面的图像。 Python包 pyhh软件包包括Hodgkin-Huxley模型和其他用于组织模拟数据的工具。 模拟步骤 创建模型单元并根据需要自定义其属性 创建刺激波形(一个numpy数组) 通过给它建模您创建的波形来创建仿真 绘制刺激的各种特性 用法示例 # customize a neuron model if desired model = pyhh . HHModel () model . gNa = 100 # typically 120 model . gK = 5 # typically 36 model . EK = - 35 # typically
资源推荐
资源详情
资源评论
收起资源包目录
 pyHH-master.zip (16个子文件)
pyHH-master.zip (16个子文件)  pyHH-master
pyHH-master  tests
tests  demo.py 2KB
demo.py 2KB demo.png 170KB
demo.png 170KB LICENSE 1KB
LICENSE 1KB src
src  pyhh
pyhh  models.py 2KB
models.py 2KB __init__.py 71B
__init__.py 71B simulations.py 1KB
simulations.py 1KB dev
dev  concept1.py 2KB
concept1.py 2KB figures
figures  320px-Hodgkin-Huxley.svg.png 5KB
320px-Hodgkin-Huxley.svg.png 5KB 800px-Hodgkin-Huxley.svg.png 16KB
800px-Hodgkin-Huxley.svg.png 16KB concept2.py 4KB
concept2.py 4KB concept3.py 3KB
concept3.py 3KB concept4.png 44KB
concept4.png 44KB concept4.py 3KB
concept4.py 3KB concept3.png 90KB
concept3.png 90KB .gitignore 1KB
.gitignore 1KB README.md 4KB
README.md 4KB共 16 条
- 1
资源评论

 咬一口咔嚓2021-12-28有没有matlab可以用的代码
咬一口咔嚓2021-12-28有没有matlab可以用的代码
weixin_38747818
- 粉丝: 9
- 资源: 894
上传资源 快速赚钱
 我的内容管理
展开
我的内容管理
展开
 我的资源
快来上传第一个资源
我的资源
快来上传第一个资源
 我的收益 登录查看自己的收益
我的收益 登录查看自己的收益 我的积分
登录查看自己的积分
我的积分
登录查看自己的积分
 我的C币
登录后查看C币余额
我的C币
登录后查看C币余额
 我的收藏
我的收藏  我的下载
我的下载  下载帮助
下载帮助

 前往需求广场,查看用户热搜
前往需求广场,查看用户热搜安全验证
文档复制为VIP权益,开通VIP直接复制
 信息提交成功
信息提交成功
