Progress In Electromagnetics Research, Vol. 130, 369–388, 2012
AN MR BRAIN IMAGES CLASSIFIER VIA PRINCIPAL
COMPONENT ANALYSIS AND KERNEL SUPPORT
VECTOR MACHINE
Y. Zhang
*
and L. Wu
School of Information Science and Engineering, Southeast University,
Nanjing, China
Abstract—Automated and accurate classification of MR brain images
is extremely important for medical analysis and interpretation. Over
the last decade numerous methods have already been proposed.
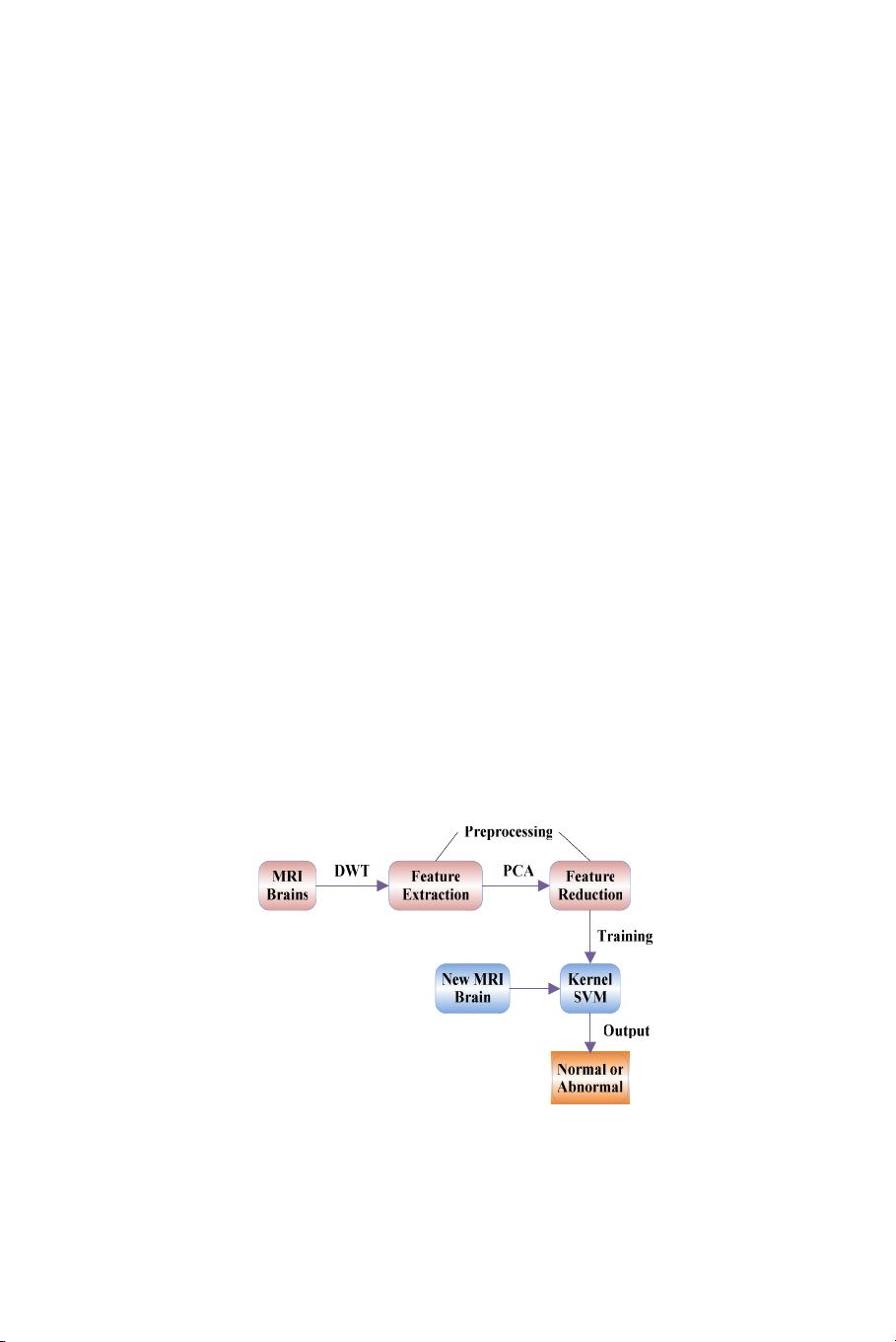
In this paper, we presented a novel method to classify a given
MR brain image as normal or abnormal. The proposed method
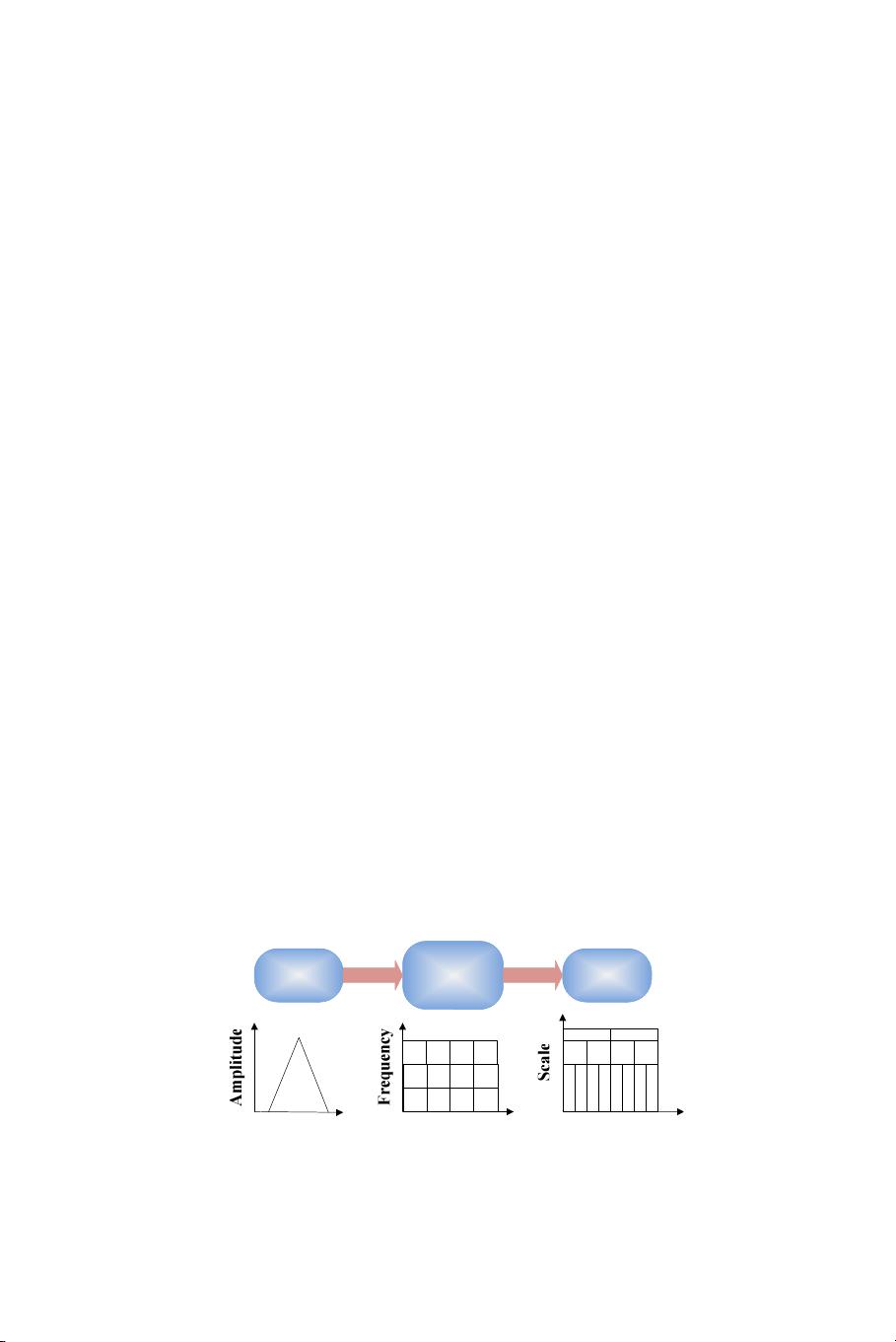
first employed wavelet transform to extract features from images,
followed by applying principle component analysis (PCA) to reduce
the dimensions of features. The reduced features were submitted
to a kernel support vector machine (KSVM). The strategy of K-
fold stratified cross validation was used to enhance generalization of
KSVM. We chose seven common brain diseases (glioma, meningioma,
Alzheimer’s disease, Alzheimer’s disease plus visual agnosia, Pick’s
disease, sarcoma, and Huntington’s disease) as abnormal brains, and
collected 160 MR brain images (20 normal and 140 abnormal) from
Harvard Medical School website. We performed our proposed methods
with four different kernels, and found that the GRB kernel achieves
the highest classification accuracy as 99.38%. The LIN, HPOL, and
IPOL kernel achieves 95%, 96.88%, and 98.12%, respectively. We also
compared our method to those from literatures in the last decade,
and the results showed our DWT+PCA+KSVM with GRB kernel
still achieved the best accurate classification results. The averaged
processing time for a 256 ×256 size image on a laptop of P4 IBM with
3 GHz processor and 2 GB RAM is 0.0448 s. From the experimental
data, our method was effective and rapid. It could be applied to the
field of MR brain image classification and can assist the doctors to
diagnose where a patient is normal or abnormal to certain degrees.
Received 14 June 2012, Accepted 23 July 2012, Scheduled 19 August 2012