
the only viable alternative is ribosomal depletion. Another
consideration is whether to generate strand-preserving li-
braries. The first generation of Illumina-based RNA-seq
used random hexamer priming to reverse-transcribe
poly(A)-selected mRNA. This methodology did not retain
information contained on the DNA strand that is actually
expressed [1] and therefore complicates the analysis and
quantification of antisense or overlapping transcripts. Sev-
eral strand-specific protocols [2], such as the widely used
dUTP method, extend the original protocol by incorporat-
ing UTP nucleotides during the second cDNA synthesis
step, prior to adapter ligation followed by digestion of the
strand containing dUTP [3]. In all cases, the size of the
final fragments (usually less than 500 bp for Illumina) will
be crucial for proper sequencing and subsequent analysis.
Furthermore, sequencing can involve single-end (SE) or
paired-end (PE) reads, although the latter is preferable for
de novo transcript discovery or isoform expression ana-
lysis [4, 5]. Similarly, longer reads improve mappability
and transcript identification [5, 6]. The best sequencing
option depends on the analysis goals. The cheaper, short
SE reads are normally sufficient for studies of gene expres-
sion levels in well-annotated organisms, whereas longer
and PE reads are preferable to characterize poorly anno-
tated transcriptomes.
Another important factor is sequencing depth or li-
brary size, which is the number of sequenced reads for a
given sample. More transcripts will be detected and their
quantification will be more precise as the sample is se-
quenced to a deeper level [1]. Nevertheless, optimal se-
quencing depth again depends on the aims of the
experiment. While some authors will argue that as few
as five million mapped reads are sufficient to quantify
accurately medium to highly expressed genes in most
eukaryotic transcriptomes, others will sequence up to
100 million reads to quantify precisely genes and tran-
scripts that have low expression levels [7]. When study-
ing single cells, which have limited sample complexity,
quantification is often carried out with just one million
reads but may be done reliably for highly expressed
genes with as few as 50,000 reads [8]; even 20,000 reads
have been used to differentiate cell types in splenic tissue
[9]. Moreover, optimal library size depends on the com-
plexity of the targeted transcriptome. Experimental results
suggest that deep sequencing improves quantification and
identification but might also result in the detection of
transcriptional noise and off-target transcripts [10]. Satur-
ation curves can be used to assess the improvement in
transcriptome coverage to be expected at a given sequen-
cing depth [10].
Finally, a crucial design factor is the number of repli-
cates. The number of replicates that should be included in
a RNA-seq experiment depends on both the amount of
technical variability in the RNA-seq procedures and the
biological variability of the system under study, as well as
on the desired statistical power (that is, the capacity for
detecting statistically significant differences in gene ex-
pression between experimental groups). These two aspects
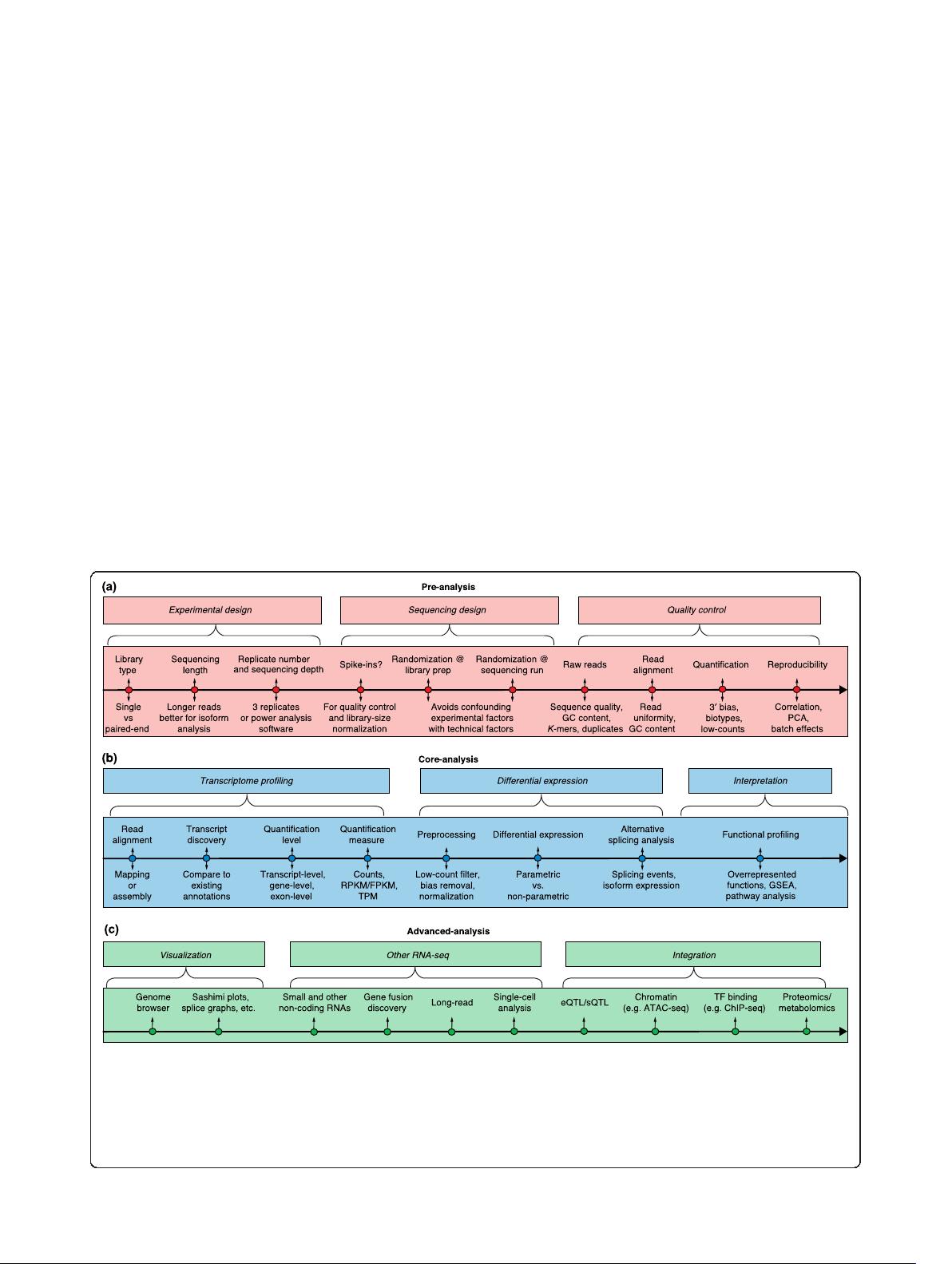
are part of power analysis calculations (Fig. 1a; Box 1).
The adequa te planning of sequencing experiments so
as to avoid technical biases is as important as good
Box 1. Number of replicates
Three factors determine the number of replicates required in a
RNA-seq experiment. The first factor is the variability in the
measurements, which is influenced by the technical noise and
the biological variation. While reproducibility in RNA-seq is usually
high at the level of sequencing [1, 45], other steps such as RNA
extraction and library preparation are noisier and may introduce
biases in the data that can be minimized by adopting good
experimental procedures (Box 2). Biological variability is particular
to each experimental system and is harder to control [189].
Nevertheless, biological replication is required if inference on the
population is to be ma de, with three replicates being the minimum
for any inferential analysis. For a proper statistical power analysis,
estimates of the within-group variance and gene expression levels
are required. This information is typically not available beforeha nd
but can be obtained from similar experiments. The exact power will
depend on the method used for differential expression analysis,
and software packages exist that provide a theoretical estimate of
power over a range of variables, given the within-group variance of
the samples, which is intrinsic to the experiment [190, 191]. Table 1
shows an example of statistical power calculations over a range of
fold-changes (or effect sizes) and number of replicates in a human
blood RNA-seq sample sequenced at 30 million mapped reads. It
should be noted that these estimates apply to the average gene
expression level, but as dynamic ranges in RNA-seq data are large,
the probability that highly expressed genes will be detected as
differentially expressed is greater than that for low-count genes
[192]. For methods that return a false discovery rate (FDR), the
proportion of genes that are highly expressed out of the total set
of genes being tested will also influence the power of detection
after multiple testing correction [193]. Filtering out genes that are
expressed at low levels prior to differential expression analysis
reduces the severity of the correction and may improve the power
of detection [20]. Increasing sequencing depth also can improve
statistical power for lowly expressed genes [10, 194], and for any
given sample there exists a level of sequencing at which power
improvement is best achieved by increasing the number of
replicates [195]. Tools such as Scotty are available to calculate the
best trade-off between sequencing depth and replicate number
given some budgetary constraints [191].
Conesa et al. Genome Biology (2016) 17:13 Page 3 of 19




 我的内容管理
展开
我的内容管理
展开
 我的资源
快来上传第一个资源
我的资源
快来上传第一个资源
 我的收益 登录查看自己的收益
我的收益 登录查看自己的收益 我的积分
登录查看自己的积分
我的积分
登录查看自己的积分
 我的C币
登录后查看C币余额
我的C币
登录后查看C币余额
 我的收藏
我的收藏  我的下载
我的下载  下载帮助
下载帮助 
 前往需求广场,查看用户热搜
前往需求广场,查看用户热搜 信息提交成功
信息提交成功

评论0