没有合适的资源?快使用搜索试试~ 我知道了~
Spiking Neural Networks and Bio-Inspired Supervised Deep Learnin
需积分: 0 0 下载量 116 浏览量
2023-08-02
11:28:27
上传
评论
收藏 1.04MB PDF 举报
温馨提示

试读
31页
SNN+生物启发式监督深度学习 内容简介: 长期以来,生物学和神经科学领域一直是计算机科学家在发展人工智能(AI)技术方面的重要灵感来源。本综述旨在全面回顾近期用于AI的受生物启发的方法。在介绍生物神经元计算和突触可塑性的主要原理后,全面介绍了脉冲神经网络(SNN)模型,并强调与SNN训练相关的主要挑战,传统的反向传播优化方法不直接适用于SNN。因此,讨论了最近的生物启发式训练方法,这些方法被视为传统网络和脉冲网络中替代反向传播的选择。生物启发的深度学习(BIDL)方法旨在推进当前模型的计算能力和生物可信性。
资源推荐
资源详情
资源评论

Spiking Neural Networks and Bio-Inspired Supervised Deep Learning: A Survey
GABRIELE LAGANI
∗
, FABRIZIO FALCHI, CLAUDIO GENNARO, and GIUSEPPE AMATO, ISTI-CNR,
Italy
For a long time, biology and neuroscience elds have been a great source of inspiration for computer scientists, towards the development
of Articial Intelligence (AI) technologies. This survey aims at providing a comprehensive review of recent biologically-inspired
approaches for AI. After introducing the main principles of computation and synaptic plasticity in biological neurons, we provide a
thorough presentation of Spiking Neural Network (SNN) models, and we highlight the main challenges related to SNN training, where
traditional backprop-based optimization is not directly applicable. Therefore, we discuss recent bio-inspired training methods, which
pose themselves as alternatives to backprop, both for traditional and spiking networks. Bio-Inspired Deep Learning (BIDL) approaches
towards advancing the computational capabilities and biological plausibility of current models.
CCS Concepts: • Computing methodologies → Bio-inspired approaches; Bio-inspired approaches.
Additional Key Words and Phrases: Bio-Inspired, Hebbian, Deep Learning, Neural Networks, Spiking
ACM Reference Format:
Gabriele Lagani, Fabrizio Falchi, Claudio Gennaro, and Giuseppe Amato. 2018. Spiking Neural Networks and Bio-Inspired Supervised
Deep Learning: A Survey. 1, 1 (August 2018), 31 pages. https://doi.org/XXXXXXX.XXXXXXX
1 INTRODUCTION
Spiking Neural Networks (SNN) have recently emerged as a biologically inspired alternative to traditional Deep Learning
(DL) models, towards addressing the current limitations of Deep Neural Networks (DNNs) in terms of ecological impact
[
9
]. Indeed, biological brains exhibit extraordinary capabilities in terms of energy eciency, supporting advanced
cognitive functions while consuming only 20W [
89
]. It is believed that the key to the energy ecient computation
of biological neurons lies in the particular coding paradigm based on short pulses, or spikes [
61
]. SNN models aim at
simulating the behavior of biological neurons more realistically, compared to traditional DNNs. As a result, SNNs are
well suited for energy-ecient implementations in neuromorphic [
84
,
174
,
186
,
190
,
229
] or biological [
92
,
111
,
176
]
hardware. This makes SNNs a promising direction toward energy-ecient DL.
Unfortunately, training SNNs is not trivial, as traditional optimization based on the backpropagation algorithm
(backprop) is not directly applicable [
165
]. In fact, the biological plausibility of backprop – the workhorse of DL – is
questioned by neuroscientists [
73
,
113
,
130
,
157
,
172
]. Therefore, researchers took again inspiration from biology, in
order to nd new learning solutions as alternatives to backprop. The goal was not only to address the problem of
SNN training [
33
,
148
], but also to discover novel approaches to the learning problem [
77
,
139
,
182
], and possibly more
data ecient strategies [
69
,
90
,
105
–
107
,
110
]. In fact, another limitation of current DL solutions is the requirement of
∗
Corresp.
Authors’ address: Gabriele Lagani, gabriele.lagani@isti.cnr.it; Fabrizio Falchi, fabrizio.falchi@isti.cnr.it; Claudio Gennaro, claudio.gennaro@isti.cnr.it;
Giuseppe Amato, giuseppe.amato@isti.cnr.it, ISTI-CNR, Pisa, Italy, 56124.
Permission to make digital or hard copies of all or part of this work for personal or classroom use is granted without fee provided that copies are not
made or distributed for prot or commercial advantage and that copies bear this notice and the full citation on the rst page. Copyrights for components
of this work owned by others than ACM must be honored. Abstracting with credit is permitted. To copy otherwise, or republish, to post on servers or to
redistribute to lists, requires prior specic permission and/or a fee. Request permissions from permissions@acm.org.
© 2018 Association for Computing Machinery.
Manuscript submitted to ACM
Manuscript submitted to ACM 1
arXiv:2307.16235v1 [cs.NE] 30 Jul 2023

2 Lagani, et al.
Fig. 1. A schematic view of the topics of Bio-Inspired Deep Learning (BIDL) addressed in this work. We discuss SNN models, which
aim at providing a biologically faithful simulation of real neurons, and they find applications in the context of energy-eicient
biological and neuromorphic computing. Learning strategies for SNNs include Spike Time Dependent Plasticity (STDP), surrogate
gradient strategies for adapting backprop optimization to the case of SNNs, etc. We also discuss training algorithms that pose
themselves as alternatives to backprop. Some approaches aim at approximating backprop using only biologically plausible synaptic
updates, while others, rather than simply approximating backprop, consider dierent perspectives, such as forward propagation
approaches, reservoir computing, etc.
large amounts of training data. On the other hand, biological brains also show interesting properties in terms of data
eciency, being able to infer new knowledge and easily generalize from little experience [112].
In this survey, we illustrate biologically realistic SNN models of neural computation, and we discuss spike coding
strategies, spiking models of synaptic plasticity, and the training challenges and application potentials of this line of
research. We also consider the learning problem in more detail, illustrating recently proposed methodologies for SNN
and DNN training, which pose themselves as alternatives to traditional backprop-based training.
One of the goals of this survey is to provide a comprehensive review of the methods coming from biological
inspiration, and their potential impact on current DL technologies. The eld of Bio-Inspired Deep Learning (BIDL)
observes the convergence of a broad spectrum of ideas, coming from the computer science and neuroscience elds;
hence, one of the goals of this work is to highlight the connections between these two viewpoints. Fig. 1 provides a
schematic visualization of the topics addressed in this work.
This document can be of interest both to novel readers who approach themes in the BIDL eld for the rst time,
as well as a reference for researchers already familiar with these topics. Moreover, this document does not require
prerequisite knowledge in the neuroscience domain, but the necessary background is provided where needed. At the
same time, neuroscientists that are curious about the engineering aspects behind AI architectures could also nd this
document an interesting resource.
Manuscript submitted to ACM

Bio-Inspired Deep Learning 3
This survey is organized as follows:
• Section 2 discusses related surveys in the BIDL eld.
• Section 3 briey introduces some background about biological neurons and synaptic plasticity models.
•
Section 4 discusses biologically detailed models of neural computation based on SNNs, highlights the challenges
of training such models, but also their technological potential for energy-ecient biological and neuromorphic
computing.
• Section 5 explores alternative DNN training algorithms which do not require backprop.
•
Finally, we present our conclusions in Section 6, outlining open challenges and possible future research directions.
Moreover, in our companion paper [
5
] we discuss biological models of synaptic plasticity in greater detail, and we
highlight the relationships between such models and unsupervised pattern discovery mechanisms in neural networks,
showing the connections between neuroscientic principles and emerging cognitive behavior.
2 RELATED SURVEYS
The eld of BIDL has been the subject of increasing attention recently, and it has been reviewed in a number of recent
contributions from dierent perspectives. In particular, there are two perspectives from which the eld is approached:
the neuroscience and the computer science/engineering viewpoints. Related contributions so far have been more tied to
one of these viewpoints, or they have tackled specic aspects of this eld. In this perspective, the goal of our contribution
is to provide a broader perspective on the various concepts that emerge in the eld of BIDL, and to highlight the
connections between the neuroscience viewpoint, and the computer science/engineering aspects.
A number of recent reviews [
73
,
113
,
130
,
172
] address the eld of BIDL from a high-level perspective, discussing
the mutual benets that exploration in biologically grounded mechanisms behind intelligence could bring both to the
neuroscience and computer science elds. The authors also suggest possible specic directions of exploration where
biological inspiration could play a crucial role.
A general background about biologically grounded neural system modeling can be found in the book from Gerstner
and Kistler [
61
], where a thorough introduction to SNN models is provided, as well as to Hebbian and Spike Time
Dependent Plasticity (STDP) models. A review more specically focused on reward-modulated Hebbian learning
approaches can be found in [
202
], while recent SNN developments and applications are surveyed in [
151
,
160
,
196
].
The discussions in [
45
,
148
] focus more specically on backprop alternatives for SNNs. Finally, various recent surveys
provide a thorough analysis of the eld of neuromorphic hardware [84, 174, 186, 190, 229].
Compared to previous surveys, we provide signicant contributions in the following directions:
•
We emphasize the connections between low-level learning mechanisms and high-level abstractions, as opposed
to works more focused on the high-level perspectives [73, 113, 130, 172];
•
We develop a comprehensive viewpoint involving bio-inspired techniques ranging from synaptic plasticity
models for spiking neurons, to supervised or reward-driven backprop alternatives, while other contributions
only focus on backprop extensions for SNNs [45, 148] or reward-driven approaches [202];
•
We highlight the connections between traditional and spike-based models, showing the interplay between the
two domains, and the potentials of SNN models for energy-ecient neuromorphic and biological computing,
compared to works that focus more specically on the low-level mechanisms of spike-based computation
[151, 160, 196].
Manuscript submitted to ACM

4 Lagani, et al.
3 BIOLOGICAL NEURONS AND SYNAPTIC PLASTICITY
This Section is devoted to introducing the fundamental aspects of neuron biology, and the mechanisms of synaptic
plasticity underlying the learning behavior of biological brains. From this it will be possible to draw relationships
between the computational learning mechanisms that we are going to discuss in the following, and the biological
substrate that supports such mechanisms.
In the following, we start with an introduction on neural cell biology in subsection 3.1, then we move to synaptic
plasticity models based on the Hebbian principle in subsection 3.2, highlighting the relationships between such models
and emerging unsupervised pattern discovery mechanisms (such as clustering and subspace learning).
3.1 Background on Neural Cells
Neural cells are characterized by a central body, the soma, which receives electric input stimuli through ramied
connections called dendrites, and emit signals through an output connection called axon [
13
,
36
,
124
,
188
,
213
]. The axon
is connected to other neurons’ dendrites by chemical couplings called synapses. The strength of synaptic couplings is
plastic, and its change determines the learning properties of neurons. The main aspects of plasticity are strengthening of
synaptic ecacy, a.k.a. Long Term Potentiation (LTP), or weakening, a.k.a. Long Term Depression (LTD) [
14
,
61
,
188
,
213
].
Upon receiving input stimuli, an electric potential is accumulated on the neuronal membrane (i.e. the membrane potential).
When the membrane potential exceeds a threshold, an output stimulus is triggered, in terms of a pulse-shaped (a spike)
electric signal propagated on the axon [61].
We can distinguish two main groups of neural cells: pyramidal and non-pyramidal [213].
Pyramidal cells represent the main computing unit in the brain. As the name suggest, they are characterized by a
pyramid-like shape, from which two types of dendritic trees branch out: apical and basal. The former extends from the
tip of the pyramid, and extend through cortical layers, while the latter extend mainly toward neighboring cells in the
same region. From the base of the pyramid, the axon of the neural cell originates. I then branches into a projection axon,
which can extend towards deeper cortical layers, and several axon collaterals, which can either be local, i.e. extending
for a short distance towards neighboring neurons, or they can extend for longer distances either in the same layer or
towards neighboring layers.
Non-pyramidal cells are instead a broad category comprising a variety of neural cell types [
194
], but all sharing
some common features such as the presence of a central body with a smaller size compared to pyramidal cells, the
local connectivity patterns (i.e. short-distance), and the mainly inhibitory role of such neurons. A dense dendritic tree
originates from the cell body, as well as an axon which tends to branch into multiple ramications. Due to the local
nature of non-pyramidal cell connectivity, which typically tend to target neighboring pyramidal cells, thus routing
information in the local neighborhood, these neurons are also often referred to as interneurons. Interneurons play an
important role in inhibitory interaction and shunting inhibition [
100
] between neurons. The mechanisms of inhibitory
lateral interaction, mediated by non-pyramidal cells, are essential to achieve decorrelation in neural activity. In the
following we will discuss how these mechanisms, coupled with appropriate synaptic plasticity models, allow neural
circuits to implement unsupervised algorithms for automatic pattern discovery, thus showing an interesting mapping
between biological circuits and learning mechanisms.
Manuscript submitted to ACM
Bio-Inspired Deep Learning 5
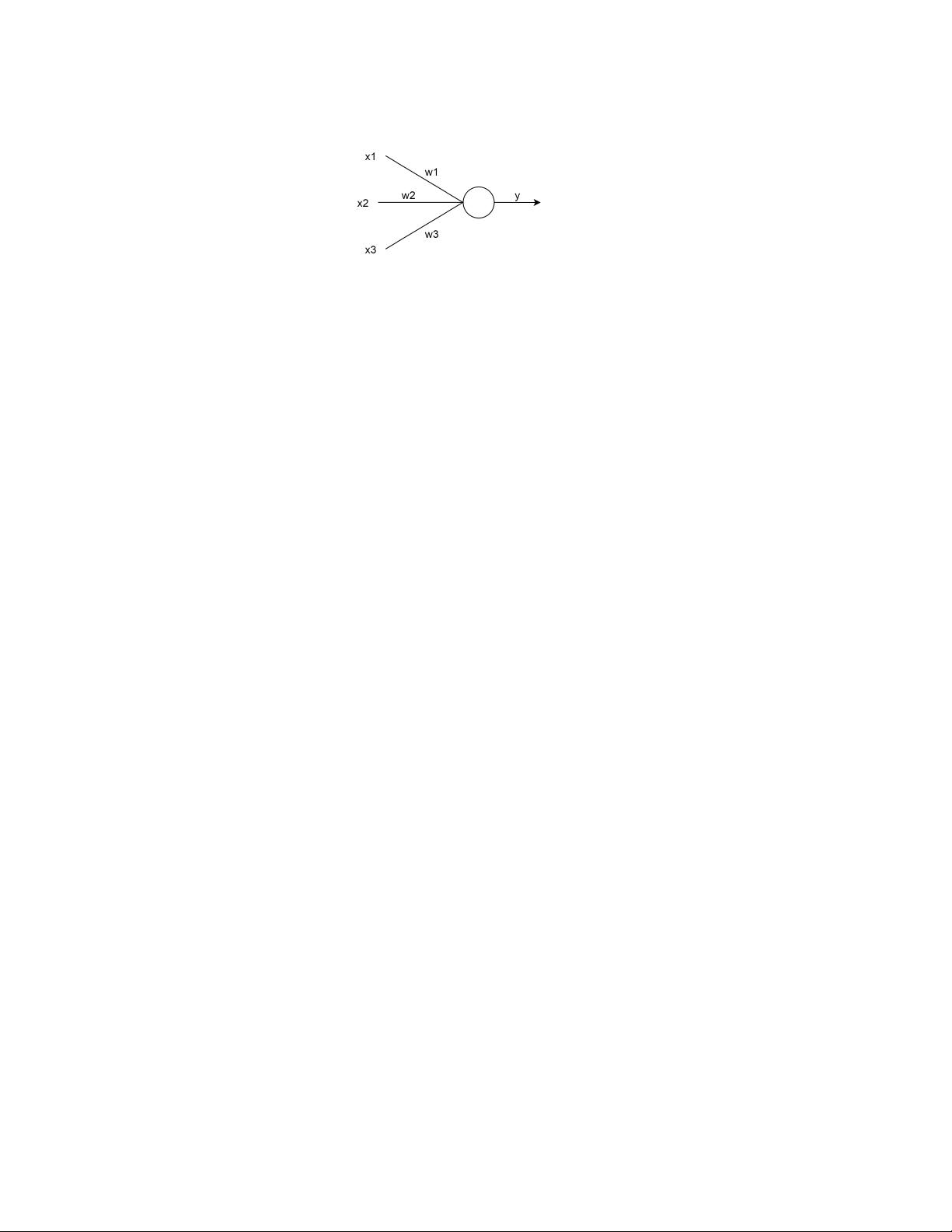
Fig. 2. Representation of a neuron model with weights
𝑤
1
, 𝑤
2
, 𝑤
3
, taking inputs
𝑥
1
, 𝑥
2
, 𝑥
3
, and computing output
𝑦 = 𝑓 (
Í
3
𝑖=1
𝑥
𝑖
𝑤
𝑖
)
3.2 Biological Models of Synaptic Plasticity
In the following we will consider a neuron whose synaptic weights are described by a vector w. The neuron takes as
input a vector x, and produces an output
𝑦(
x
,
w
) = 𝑓 (
x
𝑇
w
)
(Fig. 2), where
𝑓
is the activation function (optionally, a
bias term can be implicitly modeled as a synapse connected to a constant input). In the following, we use boldface
fonts to denote vectors, and normal fonts to denote scalars. In neuroscientic terms, input values x are also termed
pre-synaptic activations, while the output
𝑦
is termed post-synaptic activation. In order to draw a relationship between
this abstract neuron model, and the biological neural cell structure described in the previous subsection, we can imagine
that the output value of the abstract neuron model represents the rate of output spikes generated by the neuron due to
the input stimuli, hence we can talk about a rate-coded neuron model [
61
]. Similarly, the input value on a given synapse
corresponds to the rate of input spikes delivered through that synapse.
In order to model synaptic plasticity, neuroscientists propose the Hebbian principle [
61
,
74
]: "re together, wire
together". According to this principle, the synaptic coupling between two neurons is reinforced when the two neurons
are simultaneously active [
61
,
74
]. Mathematically, this learning rule, in its simplest "vanilla" formulation, can be
expressed as:
Δ𝑤
𝑖
= 𝜂 𝑦 𝑥
𝑖
(1)
where
𝜂
is the learning rate, and the subscript
𝑖
refers to the i-th input/synapse. The eect of this learning rule
is essentially to consolidate correlated activations between neural inputs and outputs, by reinforcing the synaptic
couplings, so that, if a similar input will be observed again in the future, a similar response will likely be elicited from
the neuron.
The problem with Eq. 1 is that it is unstable, as repeated stimulation can induce the weights to grow unbounded.
This issue can be counteracted by iteratively normalizing the weight vector [
152
], or by adding a weight decay term
𝛾 (w, x) to the learning rule:
Δ𝑤
𝑖
= 𝜂 𝑦 𝑥
𝑖
− 𝛾 (w, x) (2)
In particular, with an appropriate choice for this term, i.e.
𝛾 (
w
,
x
) = 𝜂 𝑦(
w
,
x
) 𝑤
𝑖
, we obtain a learning rule that has
been widely applied to the context of competitive learning [67, 96, 178]:
Δ𝑤
𝑖
= 𝜂 𝑦 𝑥
𝑖
− 𝜂 𝑦 𝑤
𝑖
= 𝜂 𝑦 (𝑥
𝑖
− 𝑤
𝑖
) (3)
It can be shown that learning rule in Eq. 1 induces the weight vector of a neuron to align towards the principal
component of the data distribution it is trained on [
61
,
152
], while Eq. 3 pushes the weight vector towards the centroid of
the cluster formed by the data points [
67
,
74
]. This shows surprising connections between biological models of synaptic
plasticity and computer science aspects related to data engineering and unsupervised pattern discovery mechanisms,
specically Principal Component Analysis (PCA) and clustering.
Manuscript submitted to ACM
剩余30页未读,继续阅读
资源评论

学术菜鸟小晨
- 粉丝: 1w+
- 资源: 4953
上传资源 快速赚钱
 我的内容管理
展开
我的内容管理
展开
 我的资源
快来上传第一个资源
我的资源
快来上传第一个资源
 我的收益 登录查看自己的收益
我的收益 登录查看自己的收益 我的积分
登录查看自己的积分
我的积分
登录查看自己的积分
 我的C币
登录后查看C币余额
我的C币
登录后查看C币余额
 我的收藏
我的收藏  我的下载
我的下载  下载帮助
下载帮助

 前往需求广场,查看用户热搜
前往需求广场,查看用户热搜最新资源
- 论文(最终)_20240430235101.pdf
- 基于python编写的Keras深度学习框架开发,利用卷积神经网络CNN,快速识别图片并进行分类
- 最全空间计量实证方法(空间杜宾模型和检验以及结果解释文档).txt
- 5uonly.apk
- 蓝桥杯Python组的历年真题
- 2023-04-06-项目笔记 - 第一百十九阶段 - 4.4.2.117全局变量的作用域-117 -2024.04.30
- 2023-04-06-项目笔记 - 第一百十九阶段 - 4.4.2.117全局变量的作用域-117 -2024.04.30
- 前端开发技术实验报告:内含4四实验&实验报告
- Highlight Plus v20.0.1
- 林周瑜-论文.docx
资源上传下载、课程学习等过程中有任何疑问或建议,欢迎提出宝贵意见哦~我们会及时处理!
点击此处反馈



安全验证
文档复制为VIP权益,开通VIP直接复制
 信息提交成功
信息提交成功