
Highly accurate protein structure prediction
with AlphaFold
In the format provided by the
authors and unedited
Nature | www.nature.com/nature
Supplementary information
https://doi.org/10.1038/s41586-021-03819-2
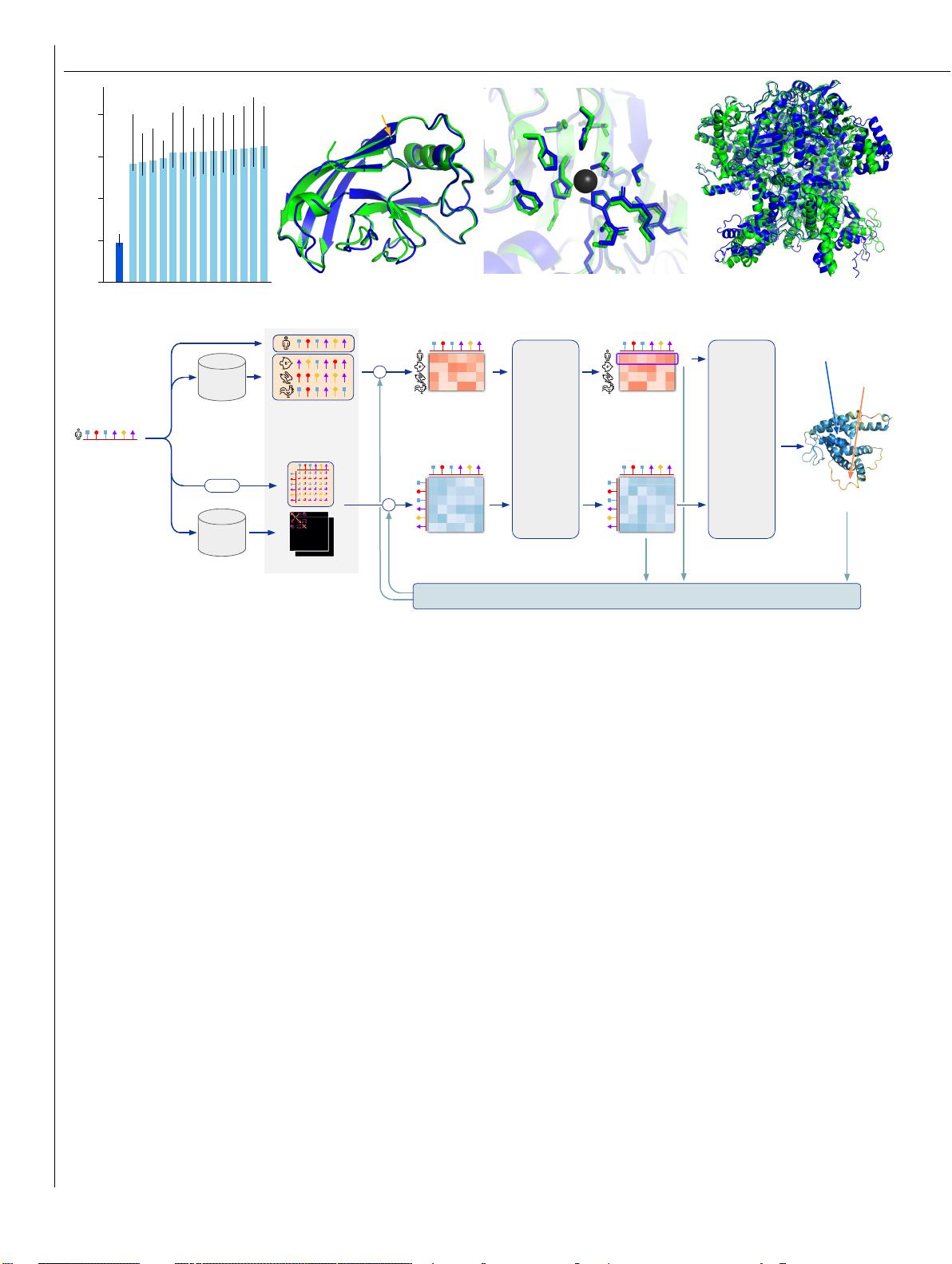
Supplementary Information for: Highly accurate
protein structure prediction with AlphaFold
John Jumper
1*+
, Richard Evans
1*
, Alexander Pritzel
1*
, Tim Green
1*
, Michael
Figurnov
1*
, Olaf Ronneberger
1*
, Kathryn Tunyasuvunakool
1*
, Russ Bates
1*
,
Augustin Žídek
1*
, Anna Potapenko
1*
, Alex Bridgland
1*
, Clemens Meyer
1*
, Simon
A A Kohl
1*
, Andrew J Ballard
1*
, Andrew Cowie
1*
, Bernardino Romera-Paredes
1*
,
Stanislav Nikolov
1*
, Rishub Jain
1*
, Jonas Adler
1
, Trevor Back
1
, Stig Petersen
1
,
David Reiman
1
, Ellen Clancy
1
, Michal Zielinski
1
, Martin Steinegger
2
, Michalina
Pacholska
1
, Tamas Berghammer
1
, Sebastian Bodenstein
1
, David Silver
1
, Oriol
Vinyals
1
, Andrew W Senior
1
, Koray Kavukcuoglu
1
, Pushmeet Kohli
1
, and Demis
Hassabis
1*+
1
DeepMind, London, UK
2
School of Biological Sciences and Artificial Intelligence Institute, Seoul National
University, Seoul, South Korea
*
These authors contributed equally
+
Corresponding authors: John Jumper, Demis Hassabis
Contents
1 Supplementary Methods 4
1.1 Notation . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 4
1.2 Data pipeline . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 5
1.2.1 Parsing . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 5
1.2.2 Genetic search . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 5
1.2.3 Template search . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 5
1.2.4 Training data . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 6
1.2.5 Filtering . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 6
1.2.6 MSA block deletion . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 6
1.2.7 MSA clustering . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 6
1.2.8 Residue cropping . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 7
1.2.9 Featurization and model inputs . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 8
1.3 Self-distillation dataset . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 9
1.4 AlphaFold Inference . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 10
1.5 Input embeddings . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 13
1.6 Evoformer blocks . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 14
1.6.1 MSA row-wise gated self-attention with pair bias . . . . . . . . . . . . . . . . . . . . 14
1
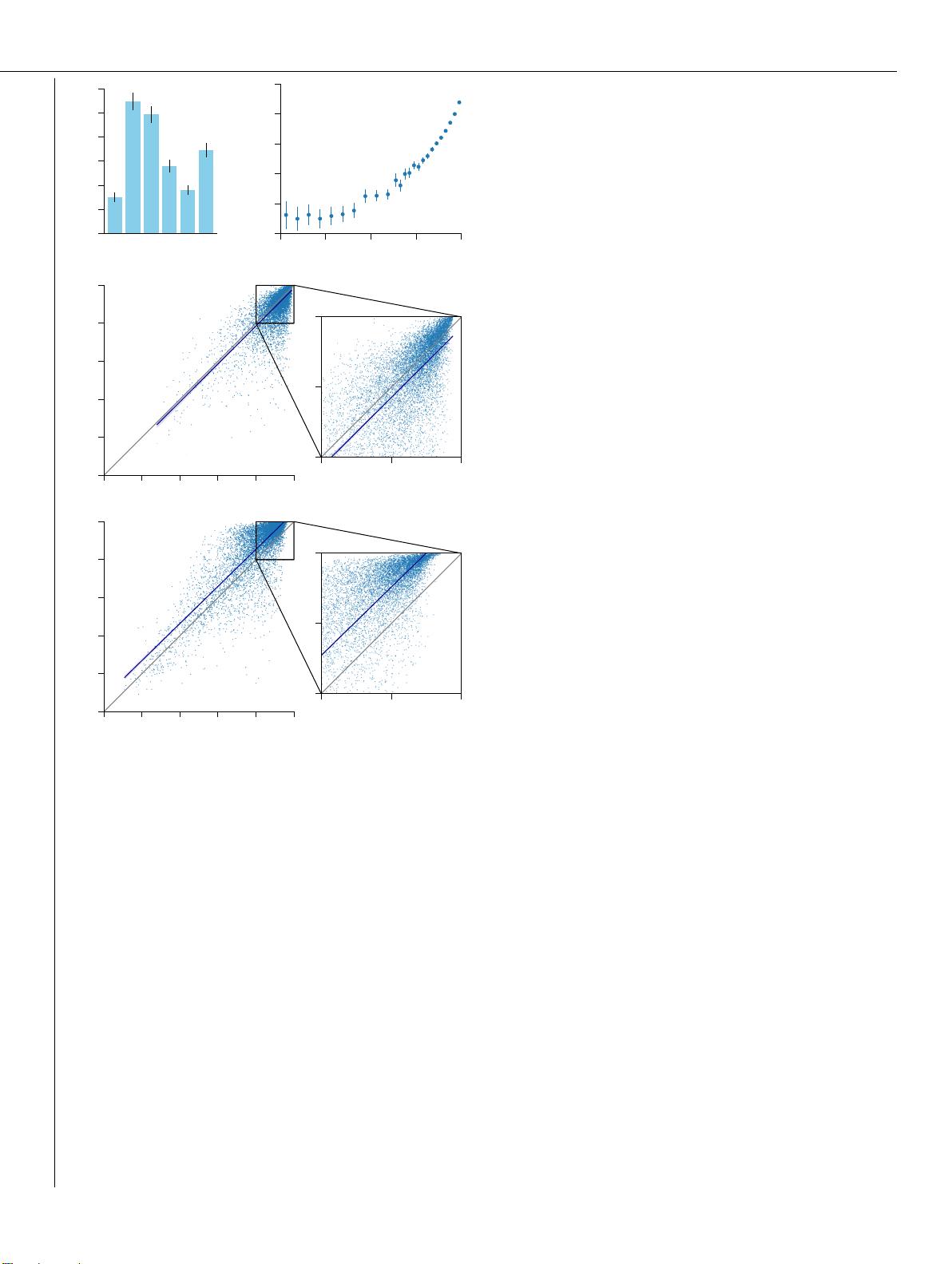
Suppl. Material for Jumper et al. (2021): Highly accurate protein structure prediction with AlphaFold 2
1.6.2 MSA column-wise gated self-attention . . . . . . . . . . . . . . . . . . . . . . . . . 15
1.6.3 MSA transition . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 16
1.6.4 Outer product mean . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 17
1.6.5 Triangular multiplicative update . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 17
1.6.6 Triangular self-attention . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 18
1.6.7 Transition in the pair stack . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 20
1.7 Additional inputs . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 20
1.7.1 Template stack . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 20
1.7.2 Unclustered MSA stack . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 21
1.8 Structure module . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 23
1.8.1 Construction of frames from ground truth atom positions . . . . . . . . . . . . . . . . 26
1.8.2 Invariant point attention (IPA) . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 26
1.8.3 Backbone update . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 28
1.8.4 Compute all atom coordinates . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 29
1.8.5 Rename symmetric ground truth atoms . . . . . . . . . . . . . . . . . . . . . . . . . 30
1.8.6 Amber relaxation . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 31
1.9 Loss functions and auxiliary heads . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 32
1.9.1 Side chain and backbone torsion angle loss . . . . . . . . . . . . . . . . . . . . . . . 32
1.9.2 Frame aligned point error (FAPE) . . . . . . . . . . . . . . . . . . . . . . . . . . . . 33
1.9.3 Chiral properties of AlphaFold and its loss . . . . . . . . . . . . . . . . . . . . . . . 34
1.9.4 Configurations with FAPE(X,Y) = 0 . . . . . . . . . . . . . . . . . . . . . . . . . . . 35
1.9.5 Metric properties of FAPE . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 36
1.9.6 Model confidence prediction (pLDDT) . . . . . . . . . . . . . . . . . . . . . . . . . 37
1.9.7 TM-score prediction . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 37
1.9.8 Distogram prediction . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 39
1.9.9 Masked MSA prediction . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 39
1.9.10 “Experimentally resolved” prediction . . . . . . . . . . . . . . . . . . . . . . . . . . 39
1.9.11 Structural violations . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 40
1.10 Recycling iterations . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 41
1.11 Training and inference details . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 43
1.11.1 Training stages . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 43
1.11.2 MSA resampling and ensembling . . . . . . . . . . . . . . . . . . . . . . . . . . . . 43
1.11.3 Optimization details . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 44
1.11.4 Parameters initialization . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 44
1.11.5 Loss clamping details . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 44
1.11.6 Dropout details . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 44
1.11.7 Evaluator setup . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 45
1.11.8 Reducing the memory consumption . . . . . . . . . . . . . . . . . . . . . . . . . . . 45
1.12 CASP14 assessment . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 46
1.12.1 Training procedure . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 46
1.12.2 Inference and scoring . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 46
1.13 Ablation studies . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 47
1.13.1 Architectural details . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 47
1.13.2 Procedure . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 49
1.13.3 Results . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 49
1.14 Network probing details . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 51
1.15 Novel fold performance . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 52
1.16 Visualization of attention . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 54
1.17 Additional results . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 56
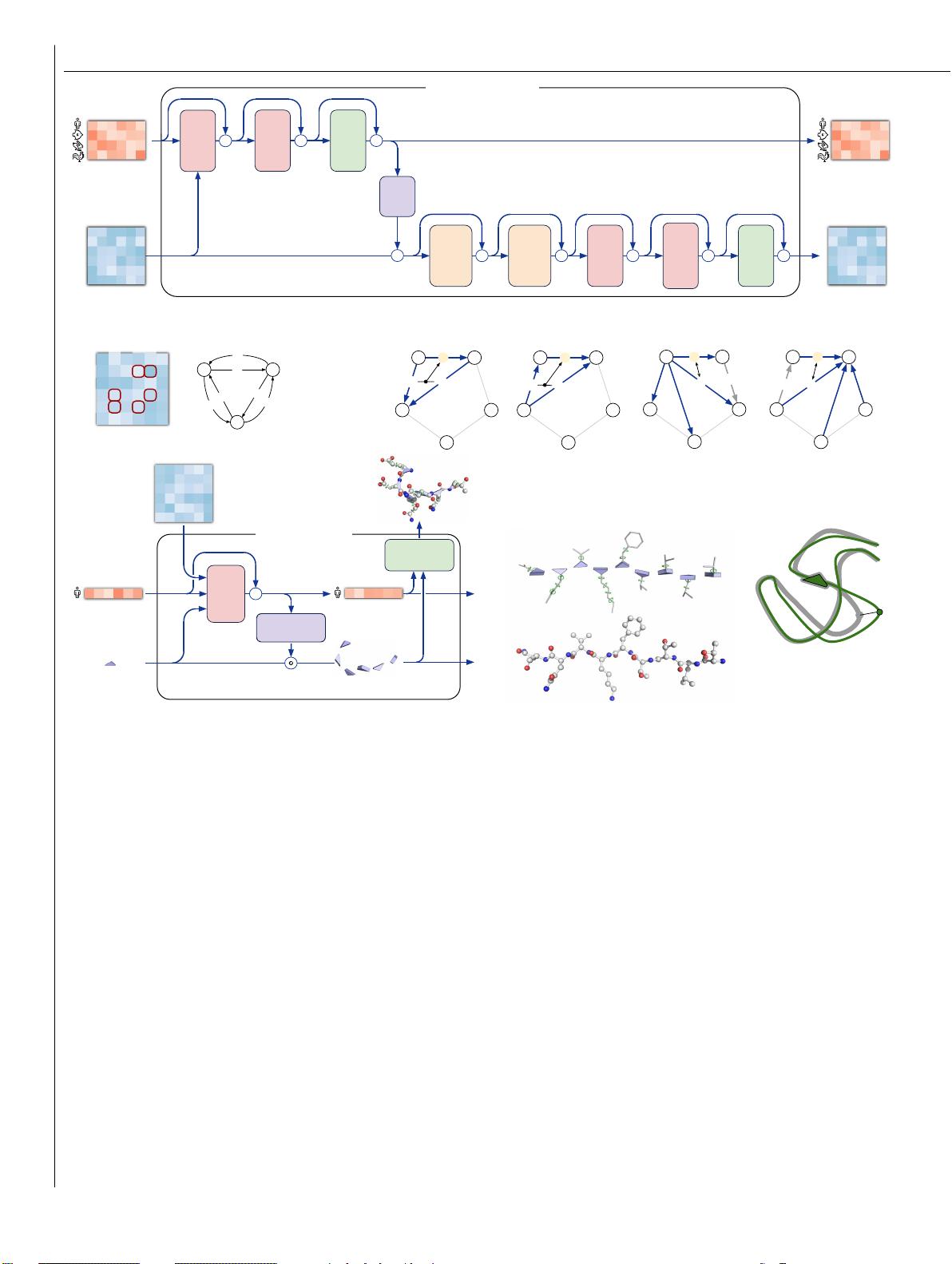
Suppl. Material for Jumper et al. (2021): Highly accurate protein structure prediction with AlphaFold 3
List of Supplementary Figures
1 Input feature embeddings . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 11
2 MSA row-wise gated self-attention with pair bias . . . . . . . . . . . . . . . . . . . . . . . . 15
3 MSA column-wise gated self-attention . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 16
4 MSA transition layer . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 16
5 Outer product mean . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 17
6 Triangular multiplicative update using “outgoing” edges . . . . . . . . . . . . . . . . . . . . 18
7 Triangular self-attention around starting node . . . . . . . . . . . . . . . . . . . . . . . . . . 19
8 Invariant Point Attention Module . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 27
9 Accuracy distribution of a model trained with dRMSD instead of the FAPE loss . . . . . . . . 36
10 Accuracy of ablations depending on the MSA depth . . . . . . . . . . . . . . . . . . . . . . . 50
11 Performance on a set of novel structures . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 53
12 Visualization of row-wise pair attention . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 54
13 Visualization of attention in the MSA along sequences . . . . . . . . . . . . . . . . . . . . . 55
14 Median all-atom RMSD
95
on the CASP14 set . . . . . . . . . . . . . . . . . . . . . . . . . . 56
15 RMSD histograms on the template-reduced recent PDB set. . . . . . . . . . . . . . . . . . . . 57
List of Supplementary Tables
1 Input features to the model . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 8
2 Rigid groups for constructing all atoms from given torsion angles . . . . . . . . . . . . . . . . 24
3 Ambiguous atom names due to 180
◦
-rotation-symmetry . . . . . . . . . . . . . . . . . . . . . 31
4 AlphaFold training protocol . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 43
5 Training protocol for CASP14 models . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 46
6 Quartiles of RMSD distributions on the template-reduced recent PDB set. . . . . . . . . . . . 57
List of Algorithms
1 MSABlockDeletion MSA block deletion . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 6
2 Inference AlphaFold Model Inference . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 12
3 InputEmbedder Embeddings for initial representations . . . . . . . . . . . . . . . . . . . . . 13
4 relpos Relative position encoding . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 13
5 one_hot One-hot encoding with nearest bin . . . . . . . . . . . . . . . . . . . . . . . . . . . 13
6 EvoformerStack Evoformer stack . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 14
7 MSARowAttentionWithPairBias MSA row-wise gated self-attention with pair bias . . . . . . 15
8 MSAColumnAttention MSA column-wise gated self-attention . . . . . . . . . . . . . . . . . . 16
9 MSATransition Transition layer in the MSA stack . . . . . . . . . . . . . . . . . . . . . . . 17
10 OuterProductMean Outer product mean . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 17
11 TriangleMultiplicationOutgoing Triangular multiplicative update using “outgoing” edges 18
12 TriangleMultiplicationIncoming Triangular multiplicative update using “incoming” edges 18
13 TriangleAttentionStartingNode Triangular gated self-attention around starting node . . . . 19
14 TriangleAttentionEndingNode Triangular gated self-attention around ending node . . . . . 20
15 PairTransition Transition layer in the pair stack . . . . . . . . . . . . . . . . . . . . . . . . 20
16 TemplatePairStack Template pair stack . . . . . . . . . . . . . . . . . . . . . . . . . . . . 21
17 TemplatePointwiseAttention Template pointwise attention . . . . . . . . . . . . . . . . . . 21
18 ExtraMsaStack Extra MSA stack . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 22
19 MSAColumnGlobalAttention MSA global column-wise gated self-attention . . . . . . . . . . 22
20 StructureModule Structure module . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 25

Suppl. Material for Jumper et al. (2021): Highly accurate protein structure prediction with AlphaFold 4
21 rigidFrom3Points Rigid from 3 points using the Gram–Schmidt process . . . . . . . . . . . 26
22 InvariantPointAttention Invariant point attention (IPA) . . . . . . . . . . . . . . . . . . . 28
23 BackboneUpdate Backbone update . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 29
24 computeAllAtomCoordinates Compute all atom coordinates . . . . . . . . . . . . . . . . . . 30
25 makeRotX Make a transformation that rotates around the x-axis . . . . . . . . . . . . . . . . . 30
26 renameSymmetricGroundTruthAtoms Rename symmetric ground truth atoms . . . . . . . . . 31
27 torsionAngleLoss Side chain and backbone torsion angle loss . . . . . . . . . . . . . . . . . 33
28 computeFAPE Compute the Frame aligned point error . . . . . . . . . . . . . . . . . . . . . . 34
29 predictPerResidueLDDT Predict model confidence pLDDT . . . . . . . . . . . . . . . . . . 37
30 RecyclingInference Generic recycling inference procedure . . . . . . . . . . . . . . . . . . 41
31 RecyclingTraining Generic recycling training procedure . . . . . . . . . . . . . . . . . . . 41
32 RecyclingEmbedder Embedding of Evoformer and Structure module outputs for recycling . . 42
1 Supplementary Methods
1.1 Notation
We denote the number of residues in the input primary sequence by N
res
(cropped during training), the number
of templates used in the model by N
templ
, the number of all available MSA sequences by N
all_seq
, the number of
clusters after MSA clustering by N
clust
, the number of sequences processed in the MSA stack by N
seq
(where
N
seq
= N
clust
+ N
templ
), and the number of unclustered MSA sequences by N
extra_seq
(after sub-sampling,
see subsubsection 1.2.7 for details). Concrete values for these parameters are given in the training details
(subsection 1.11). On the model side, we also denote the number of blocks in Evoformer-like stacks by N
block
(subsection 1.6), the number of ensembling iterations by N
ensemble
(subsubsection 1.11.2), and the number of
recycling iterations by N
cycle
(subsection 1.10).
We present architectural details in Algorithms, where we use the following conventions. We use capitalized
operator names when they encapsulate learnt parameters, e.g. we use Linear for a linear transformation with
a weights matrix W and a bias vector b, and LinearNoBias for the linear transformation without the bias
vector. Note that when we have multiple outputs from the Linear operator at the same line of an algorithm,
we imply different trainable weights for each output. We use LayerNorm for the layer normalization [85]
operating on the channel dimensions with learnable per-channel gains and biases. We also use capitalized
names for random operators, such as those related to dropout. For functions without parameters we use lower
case operator names, e.g. sigmoid, softmax, stopgrad. We use for the element-wise multiplication, ⊗
for the outer product, ⊕ for the outer sum, and a
>
b for the dot product of two vectors. Indices i, j, k
always operate on the residue dimension, indices s, t on the sequence dimension, and index h on the attention
heads dimension. The channel dimension is implicit and we type the channel-wise vectors in bold, e.g. z
ij
.
Algorithms operate on sets of such vectors, e.g. we use {z
ij
} to denote all pair representations.
In the structure module, we denote Euclidean transformations corresponding to frames by T = (R,
~
t), with
R ∈ R
3×3
for the rotation and
~
t ∈ R
3
for the translation components. We use the ◦ operator to denote
application of a transformation to an atomic position
~
x ∈ R
3
:
~
x
result
= T ◦
~
x
= (R,
~
t) ◦
~
x
= R
~
x +
~
t .
The ◦ operator also denotes composition of Euclidean transformations:
T
result
= T
1
◦ T
2
(R
result
,
~
t
result
) = (R
1
,
~
t
1
) ◦ (R
2
,
~
t
2
)
= (R
1
R
2
, R
1
~
t
2
+
~
t
1
)















