IEEE TRANSACTIONS ON MEDICAL IMAGING, VOL. 16, NO. 2, APRIL 1997 187
Multimodality Image Registration by
Maximization of Mutual Information
Frederik Maes,* Andr´e Collignon, Dirk Vandermeulen, Guy Marchal, and Paul Suetens, Member, IEEE
Abstract— A new approach to the problem of multimodality
medical image registration is proposed, using a basic concept
from information theory, mutual information (MI), or relative
entropy, as a new matching criterion. The method presented
in this paper applies MI to measure the statistical dependence
or information redundancy between the image intensities of
corresponding voxels in both images, which is assumed to be
maximal if the images are geometrically aligned. Maximization
of MI is a very general and powerful criterion, because no
assumptions are made regarding the nature of this dependence
and no limiting constraints are imposed on the image content
of the modalities involved. The accuracy of the MI criterion
is validated for rigid body registration of computed tomog-
raphy (CT), magnetic resonance (MR), and photon emission
tomography (PET) images by comparison with the stereotactic
registration solution, while robustness is evaluated with respect
to implementation issues, such as interpolation and optimization,
and image content, including partial overlap and image degra-
dation. Our results demonstrate that subvoxel accuracy with
respect to the stereotactic reference solution can be achieved
completely automatically and without any prior segmentation,
feature extraction, or other preprocessing steps which makes this
method very well suited for clinical applications.
Index Terms—Matching criterion, multimodality images, mu-
tual information, registration.
I. INTRODUCTION
T
HE geometric alignment or registration of multimodality
images is a fundamental task in numerous applications in
three-dimensional (3-D) medical image processing. Medical
diagnosis, for instance, often benefits from the complemen-
tarity of the information in images of different modalities.
In radiotherapy planning, dose calculation is based on the
computed tomography (CT) data, while tumor outlining is of-
ten better performed in the corresponding magnetic resonance
(MR) scan. For brain function analysis, MR images provide
anatomical information while functional information may be
Manuscript received February 21, 1996; revised July 23, 1996. This work
was supported in part by IBM Belgium (Academic Joint Study) and by the
Belgian National Fund for Scientific Research (NFWO) under Grants FGWO
3.0115.92, 9.0033.93 and G.3115.92. The Associate Editor responsible for
coordinating the review of this paper and recommending its publication was
N. Ayache. Asterisk indicates corresponding author.
*F. Maes is with the Laboratory for Medical Imaging Research,
Katholieke Universiteit Leuven, ESAT/ Radiologie, Universitair Ziekenhuis
Gasthuisberg, Herestraat 49, B-3000 Leuven, Belgium. He is an Aspirant
of the Belgian National Fund for Scientific Research (NFWO) (e-mail:
Frederik.Maes@uz.kuleuven.ac.be).
A. Collingnon, D. Vandermeulen, G. Marchal, and P. Suetens are with the
Laboratory for Medical Imaging Research, Katholieke Universiteit Leuven,
ESAT/Radiologie, Universitair Ziekenhuis Gasthuisberg, Herestraat 49, B-
3000 Leuven, Belgium.
Publisher Item Identifier S 0278-0062(97)02397-5.
obtained from positron emission tomography (PET) images,
etc.
The bulk of registration algorithms in medical imaging (see
[3], [16], and [23] for an overview) can be classified as being
either frame based, point landmark based, surface based, or
voxel based. Stereotactic frame-based registration is very ac-
curate, but inconvenient, and cannot be applied retrospectively,
as with any external point landmark-based method, while
anatomical point landmark-based methods are usually labor-
intensive and their accuracy depends on the accurate indication
of corresponding landmarks in all modalities. Surface-based
registration requires delineation of corresponding surfaces
in each of the images separately. But surface segmentation
algorithms are generally highly data and application dependent
and surfaces are not easily identified in functional modalities
such as PET. Voxel-based (VSB) registration methods optimize
a functional measuring the similarity of all geometrically cor-
responding voxel pairs for some feature. The main advantage
of VSB methods is that feature calculation is straightforward
or even absent when only grey-values are used, such that
the accuracy of these methods is not limited by segmentation
errors as in surface based methods.
For intramodality registration multiple VSB methods have
been proposed that optimize some global measure of the
absolute difference between image intensities of corresponding
voxels within overlapping parts or in a region of interest (ROI)
[5], [11], [19], [26]. These criteria all rely on the assumption
that the intensities of the two images are linearly correlated,
which is generally not satisfied in the case of intermodality
registration. Crosscorrelation of feature images derived from
the original image data has been applied to CT/MR matching
using geometrical features such as edges [15] and ridges [24]
or using especially designed intensity transformations [25].
But feature extraction may introduce new geometrical errors
and requires extra calculation time. Furthermore, correlation of
sparse features like edges and ridges may have a very peaked
optimum at the registration solution, but at the same time be
rather insensitive to misregistration at larger distances, as all
nonedge or nonridge voxels correlate equally well. A mul-
tiresolution optimization strategy is therefore required, which
is not necessarily a disadvantage, as it can be computationally
attractive.
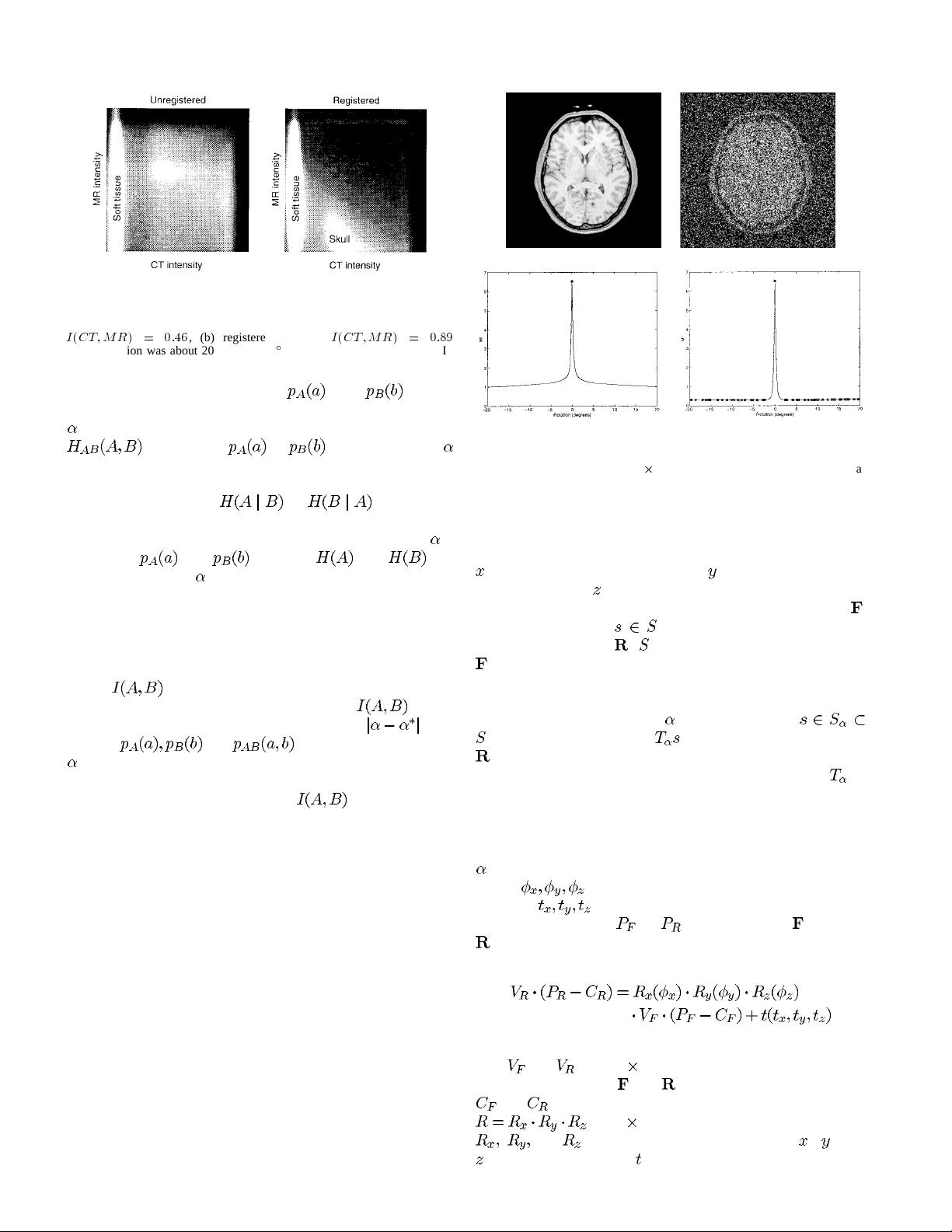
In the approach of Woods et al. [30] and Hill et al. [12],
[13], misregistration is measured by the dispersion of the
two-dimensional (2-D) histogram of the image intensities of
corresponding voxel pairs, which is assumed to be minimal
in the registered position. But the dispersion measures they
0278–0062/97$10.00 1997 IEEE